AI Unifies Cell Maps into Spatial Tissue Atlases
 A groundbreaking computational method is poised to significantly accelerate the creation of detailed spatial maps of cells within the human body, according to a recent study published in Nature Genetics. This advanced technique, known as spatial multi-omics, provides researchers with ultra-high-resolution views of tissues, enabling them to pinpoint not only which genes or proteins are active within a cell but also precisely where that activity is taking place. This spatial context is invaluable for comprehending the intricate workings of complex organs like the brain, immune tissues, and developing embryos.
A groundbreaking computational method is poised to significantly accelerate the creation of detailed spatial maps of cells within the human body, according to a recent study published in Nature Genetics. This advanced technique, known as spatial multi-omics, provides researchers with ultra-high-resolution views of tissues, enabling them to pinpoint not only which genes or proteins are active within a cell but also precisely where that activity is taking place. This spatial context is invaluable for comprehending the intricate workings of complex organs like the brain, immune tissues, and developing embryos.
However, the simultaneous capture of multiple molecular layers within a tissue remains a costly and technically demanding endeavor. David Gate, Ph.D., an assistant professor in the Ken and Ruth Davee Department of Neurology's Division of Behavioral Neurology and a co-author of the study, highlighted this challenge.
"In practice, investigators often end up with 'mosaic' datasets," explained Gate, who also leads the Abrams Research Center on Neurogenomics. "These are disparate slices or batches, each capturing only a subset of the molecular layers. They frequently come from different technologies or labs, leading to issues with batch effects and missing information."
A New Tool to Unify Complex Data
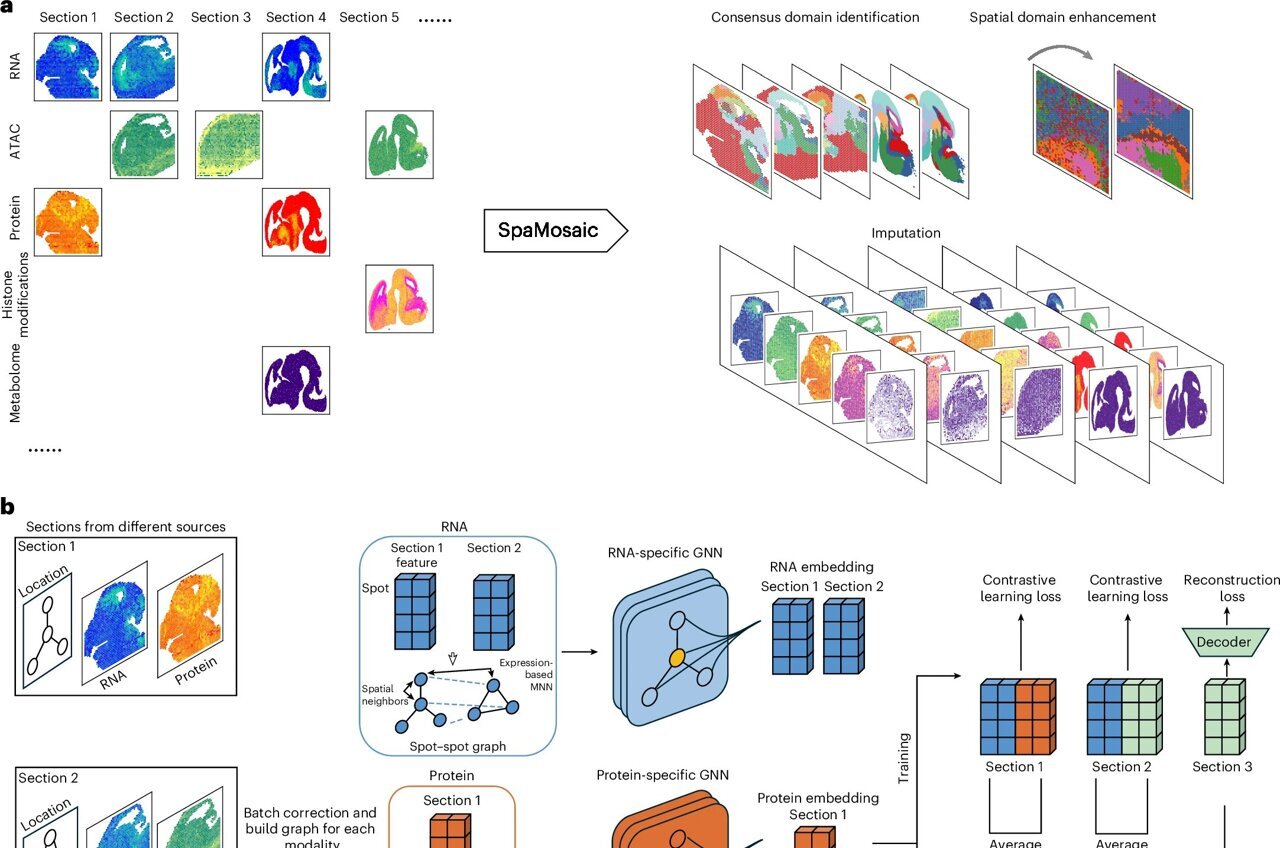
To address this escalating problem, a novel computational method called SpaMosaic has been developed. Spearheaded by a collaborative team of computational researchers, this innovative tool leverages artificial intelligence to seamlessly align and integrate diverse spatial datasets.
The creation of SpaMosaic involved a sophisticated combination of contrastive learning and graph neural networks. Contrastive learning empowers AI models to discern meaningful similarities and differences across various datasets, while graph neural networks are adept at accounting for the spatial relationships between neighboring cells. The synergy of these approaches results in a unified dataset, allowing for the simultaneous analysis of RNA, protein, chromatin accessibility, and histone modification data, even when individual datasets only measure a partial set of these molecular features.
Robust Performance Across Tissues and Species
Rigorous benchmarking experiments have demonstrated SpaMosaic's consistent superiority over existing data integration methods. The tool has proven effective on both simulated data and real-world datasets, encompassing a wide range of biological samples. These include the developing mouse brain, entire mouse embryos, and human immune tissues such as lymph nodes and tonsils. The researchers observed that SpaMosaic excelled in identifying biologically significant spatial domains—distinct regions within a tissue that share a common functional identity—regardless of whether the datasets originated from different technologies or various developmental stages.
"SpaMosaic is also highly effective at removing technical 'batch effects,' which are essentially differences arising from how samples were processed, while crucially preserving the underlying biological signals," Gate emphasized.
Predicting Unmeasured Molecular Layers
One of SpaMosaic's most remarkable innovations is its capacity to predict molecular layers that were never directly measured in the original experiments. In a comprehensive mosaic dataset derived from the mouse brain, the tool successfully inferred histone modification patterns in areas where only transcriptomic data had been collected.
"SpaMosaic effectively filled in the gaps," Gate noted. "It actually revealed stronger connections between gene activity and epigenetic regulation than the directly measured chromatin data sometimes indicated."
Implications for Tissue Atlases and Neuroscience
These findings suggest that SpaMosaic offers a powerful alternative to expensive and technically demanding experimental approaches for uncovering regulatory relationships between molecular layers. Instead of being constrained by the limitations of a single experiment, researchers can now pool and analyze data from a multitude of studies, platforms, and laboratories.
"This is a true game-changer for the construction of comprehensive multi-omics 'atlases' of tissues," Gate stated. "For neuroscience, which is our primary focus, this means we can develop far superior maps of brain development, neuroinflammation, and eventually, disease states like Alzheimer's or ALS. In these conditions, spatial relationships and multi-layer regulation are critically important. This tool accelerates discovery without requiring every lab to repeat complex, multi-modal experiments on every single sample."
Future Directions for SpaMosaic
The development team is actively pursuing the next stages of SpaMosaic's evolution. Efforts are underway to scale the tool to handle even larger and more complex datasets. Furthermore, Gate and his collaborators plan to conduct further validation studies to rigorously assess the reliability of the data predicted by the method.
"This project beautifully illustrates the power of close collaboration between computational innovators and experimental biologists," Gate concluded. "Tools like SpaMosaic are democratizing spatial multi-omics, enabling a broader range of laboratories to contribute to and benefit from the creation of large-scale tissue atlases."

Post a Comment for "AI Unifies Cell Maps into Spatial Tissue Atlases"